HIN (.hin)
HIN (.hin)
- Import fully supports HyperChem HIN files.
Background & Context
-
- MIME type: chemical/x-hin
- HyperChem HIN format.
- Used in cheminformatics applications and on the web for storing and exchanging 3D molecule models.
- Plain text tabular format.
- Stores atomic coordinates, chemical bond information, and metadata.
- Maintained by HyperCube, Inc.
Import & Export
- Import["file.hin"] imports a list of molecules from an HIN file.
- Export["file.hin",expr] exports a molecule or list of molecules to an HIN file.
- Import["file.hin",elem] imports the specified element from a molfile.
- Import["file.hin",{{elem1,elem2,…}}] imports multiple elements.
- The import format can be specified with Import["file","HIN"] or Import["file",{"HIN",elem,…}].
- Export["file.hin",mol] creates an HIN file from a molecule.
- Export["file.hin",{mol1,mol2,…}] creates an HIN file from a list of molecules.
- See the following reference pages for full general information:
-
Import, Export import from or export to a file CloudImport, CloudExport import from or export to a cloud object ImportString, ExportString import from or export to a string ImportByteArray, ExportByteArray import from or export to a byte array
Import Elements

- General Import elements:
-
"Elements" list of elements and options available in this file "Summary" summary of the file "Rules" list of rules for all available elements - Graphics elements:
-
"Graphics3D" 3D renderings of the molecules in an HIN file "StructureDiagram" 2D structure formula - Data elements:
-
"Molecule" a symbolic representation of the molecule model "Molecule", n a symbolic representation of the n 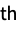 molecule
molecule"EdgeRules" connectivity data, given as an array of rules "EdgeTypes" bond types, given as an array of strings "FormalCharges" electric charges of the atoms, given as an array of integers "MassNumbers" isotope mass numbers "VertexCoordinates" 3D atomic coordinates, typically given in picometers "VertexTypes" all atoms or groups constituting the molecules, typically given as an array of chemical element abbreviations
Options

- General Import options:
-
ImageSize Automatic specifies the overall size of the graphics to display Background White specifies what background color to use ViewPoint Automatic point in space from which the 3D model is to be viewed - With the default setting "ViewPoint"->Automatic, the Wolfram Language automatically calculates the optimal viewing angle for the imported molecule geometry.
- Selecting a 3D rendering style:
-
"Rendering" "BallAndStick" specifies the visualization method - Possible settings for "Rendering" are:
-
"BallAndStick" displays atoms and bonds as a ball-and-stick model "Spacefilling" atoms shown as overlapping spheres "Wireframe" bonds rendered as lines
Examples
Basic Examples (2)
Show the Import elements available in an HIN file:
Import a molecule from an HIN file:
Show the bonds of the same molecule as a wireframe model:
History
Introduced in 2012 (9.0) | Updated in 2020 (12.1)