MoleculePlot[mol]
creates a two-dimensional structure diagram of the molecule mol.
MoleculePlot[mol,patt]
creates a diagram of mol where all atoms and bonds matching the pattern patt are highlighted.




MoleculePlot
MoleculePlot[mol]
creates a two-dimensional structure diagram of the molecule mol.
MoleculePlot[mol,patt]
creates a diagram of mol where all atoms and bonds matching the pattern patt are highlighted.
Details and Options
- MoleculePlot returns a Graphics expression.
- If the molecule does not have structure diagram coordinates, they will be generated automatically.
- mol can be a Molecule object or something that can be easily converted to one, such as a systematic chemical name, a "Chemical" entity, a BioSequence or an ExternalIdentifier.
- MoleculePlot[MoleculePattern[…]] will return a depiction of the given molecule pattern.
- Possible forms for patt include:
-
n the index of a particular atom Atom[…] a pattern for an atom Bond[…] a pattern for a bond MoleculePattern[…] a pattern for a molecule substructure {patt1,patt2,…} a list of patterns <|label1patt1,…|> an Association of labels and patterns - MoleculePlot has the same options as Graphics, with the following additions and changes: [List of all options]
-
AtomLabels Automatic labels and label placements for atoms AtomLabelStyle Automatic style to use for atom labels BondLabels None labels and label placements for bonds BondLabelStyle Automatic style to use for bond labels ColorRules Automatic a list of rules {elem1->col1,…} dictating which colors to use for atomic elements IncludeHydrogens Automatic whether to show hydrogen atoms PlotLegends None legends for highlights PlotTheme $PlotTheme overall theme for the plot - Supported plot themes include:
-
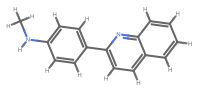
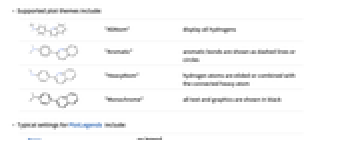
"AllAtom" display all hydrogens 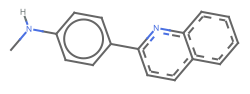
"Aromatic" aromatic bonds are shown as dashed lines or circles 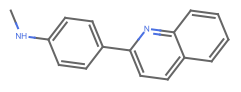
"HeavyAtom" hydrogen atoms are elided or combined with the connected heavy atom 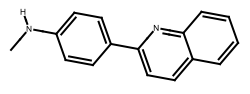
"Monochrome" all text and graphics are shown in black - Typical settings for PlotLegends include:
-
None no legend Automatic automatically determine legend {lbl1,lbl2,…} use lbl1, lbl2, … as legend labels Placed[lspec,…] specify placement for legend -
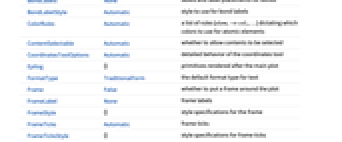
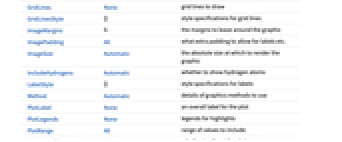
AlignmentPoint Center the default point in the graphic to align with AspectRatio Automatic ratio of height to width AtomLabels Automatic labels and label placements for atoms AtomLabelStyle Automatic style to use for atom labels Axes False whether to draw axes AxesLabel None axes labels AxesOrigin Automatic where axes should cross AxesStyle {} style specifications for the axes Background None background color for the plot BaselinePosition Automatic how to align with a surrounding text baseline BaseStyle {} base style specifications for the graphic BondLabels None labels and label placements for bonds BondLabelStyle Automatic style to use for bond labels ColorRules Automatic a list of rules {elem1->col1,…} dictating which colors to use for atomic elements ContentSelectable Automatic whether to allow contents to be selected CoordinatesToolOptions Automatic detailed behavior of the coordinates tool Epilog {} primitives rendered after the main plot FormatType TraditionalForm the default format type for text Frame False whether to put a frame around the plot FrameLabel None frame labels FrameStyle {} style specifications for the frame FrameTicks Automatic frame ticks FrameTicksStyle {} style specifications for frame ticks GridLines None grid lines to draw GridLinesStyle {} style specifications for grid lines ImageMargins 0. the margins to leave around the graphic ImagePadding All what extra padding to allow for labels etc. ImageSize Automatic the absolute size at which to render the graphic IncludeHydrogens Automatic whether to show hydrogen atoms LabelStyle {} style specifications for labels Method Automatic details of graphics methods to use PlotLabel None an overall label for the plot PlotLegends None legends for highlights PlotRange All range of values to include PlotRangeClipping False whether to clip at the plot range PlotRangePadding Automatic how much to pad the range of values PlotRegion Automatic the final display region to be filled PlotTheme $PlotTheme overall theme for the plot PreserveImageOptions Automatic whether to preserve image options when displaying new versions of the same graphic Prolog {} primitives rendered before the main plot RotateLabel True whether to rotate y labels on the frame Ticks Automatic axes ticks TicksStyle {} style specifications for axes ticks
List of all options
Examples
open all close allBasic Examples (2)
Scope (2)
Options (7)
AtomLabels (1)
AtomLabelStyle (1)
BondLabels (1)
BondLabelStyle (1)
See Also
MoleculePlot3D Molecule MoleculeRecognize AtomLabels AtomLabelStyle BondLabels BondLabelStyle
Entity Types: Chemical Protein
Function Repository: MoleculeValuePlot SmilesPlot BioSequenceMoleculePlot
Related Guides
Text
Wolfram Research (2019), MoleculePlot, Wolfram Language function, https://reference.wolfram.com/language/ref/MoleculePlot.html (updated 2021).
CMS
Wolfram Language. 2019. "MoleculePlot." Wolfram Language & System Documentation Center. Wolfram Research. Last Modified 2021. https://reference.wolfram.com/language/ref/MoleculePlot.html.
APA
Wolfram Language. (2019). MoleculePlot. Wolfram Language & System Documentation Center. Retrieved from https://reference.wolfram.com/language/ref/MoleculePlot.html
BibTeX
@misc{reference.wolfram_2025_moleculeplot, author="Wolfram Research", title="{MoleculePlot}", year="2021", howpublished="\url{https://reference.wolfram.com/language/ref/MoleculePlot.html}", note=[Accessed: 06-May-2026]}
BibLaTeX
@online{reference.wolfram_2025_moleculeplot, organization={Wolfram Research}, title={MoleculePlot}, year={2021}, url={https://reference.wolfram.com/language/ref/MoleculePlot.html}, note=[Accessed: 06-May-2026]}