NearestNeighborG[pdata,r]
estimates the nearest neighbor function ![]() at radius r in the point data pdata.
at radius r in the point data pdata.
NearestNeighborG[pproc,r]
computes ![]() for the point process pproc.
for the point process pproc.
NearestNeighborG[bdata,r]
computes ![]() for binned data bdata.
for binned data bdata.
NearestNeighborG[pspec]
generates the function ![]() that can be applied repeatedly to different radii r.
that can be applied repeatedly to different radii r.


NearestNeighborG
NearestNeighborG[pdata,r]
estimates the nearest neighbor function ![]() at radius r in the point data pdata.
at radius r in the point data pdata.
NearestNeighborG[pproc,r]
computes ![]() for the point process pproc.
for the point process pproc.
NearestNeighborG[bdata,r]
computes ![]() for binned data bdata.
for binned data bdata.
NearestNeighborG[pspec]
generates the function ![]() that can be applied repeatedly to different radii r.
that can be applied repeatedly to different radii r.
Details and Options



- NearestNeighborG is also known as nearest neighbor function and nearest neighbor distribution.
- The function
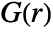 gives the probability of finding another point within distance
gives the probability of finding another point within distance 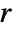 of a typical point either in pdata or in pproc.
of a typical point either in pdata or in pproc. -
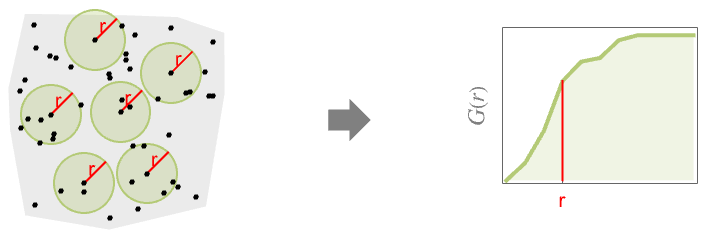
- When comparing with a Poisson point process, the results are:
-
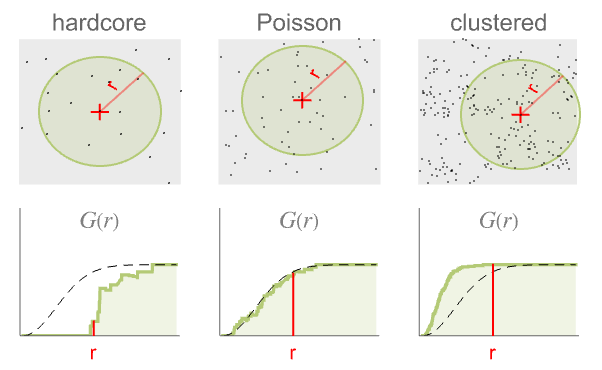
- The radius r can be a single value or a list of values. With no radius r specified, NearestNeighborG returns a PointStatisticFunction that can be used to evaluate the
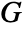 function repeatedly.
function repeatedly. - The points pdata can have the following forms:
-
{p1,p2,…} points pi GeoPosition[…],GeoPositionXYZ[…],… geographic points SpatialPointData[…] spatial point collection {pts,reg} point collection pts and observation region reg - If the observation region reg is not given, a region is automatically computed using RipleyRassonRegion.
- The point process pproc can have the following forms:
-
proc a point process proc {proc,reg} a point process proc and observation region reg - The observation region reg should be parameter free and SpatialObservationRegionQ.
- The binned data bdata is from SpatialBinnedPointData and is treated as an InhomogeneousPoissonPointProcess with a piecewise constant intensify function.
- For pdata,
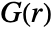 is computed by discretizing the observation region and assumes constant point intensity.
is computed by discretizing the observation region and assumes constant point intensity. - For, pproc
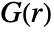 is computed by using exact formulas or by simulation to generate point data.
is computed by using exact formulas or by simulation to generate point data. - The following option can be given:
-
Method Automatic what methods to use SpatialBoundaryCorrection Automatic what boundary correction to use - The following settings can be used for SpatialBoundaryCorrection:
-
Automatic automatically determined boundary correction None no boundary correction "BorderMargin" use interior margin for observation region "Hanisch" drops points for which the distance to the nearest neighbor is greater than the distance to the boundary "KaplanMeier" SurvivalDistribution method: the point distance to its nearest neighbor is censored by its distance to the region boundary "NelsonAalen" SurvivalDistribution method: the point distance to its nearest neighbor is censored by its distance to the region boundary - The setting Method->{"Discretization"->opts} allows for adjusting the discretization method in the estimation. Here opts can be any valid options for DiscretizeRegion.
Examples
open all close allBasic Examples (3)
Estimate the nearest neighbor function at a given distance:
Estimate the nearest neighbor function within a range of distances:
Visualize the result with ListPlot:
Scope (8)
Point Data (5)
Estimate the nearest neighbor function at distance 0.1:
Obtain empirical estimates of the nearest neighbor function for a list of given distances:
Use NearestNeighborG with SpatialPointData:
Create a PointStatisticFunction for future use:
Evaluate the function at a given radius:
Estimate the nearest neighbor function without explicitly providing the observation region:
Observation region generated by Ripley–Rasson estimator:
Estimated ![]() function at distance 0.05:
function at distance 0.05:
Use NearestNeighborG with GeoPosition:
Point Processes (3)
The nearest neighbor function for PoissonPointProcess has closed form:
Visualize for fixed intensity and varying dimension:
The nearest neighbor function for a cluster process ThomasPointProcess with specified dimension:
The nearest neighbor function for a cluster process MaternPointProcess with specified dimension:
Options (2)
SpatialBoundaryCorrection (2)
The NearestNeighborG estimator without boundary correction is biased and should not be used unless with a large point set:
The default method "BorderMargin" only considers the points that are distance ![]() from the boundary:
from the boundary:
The "Hanisch" method weights each point in the observation region to make the estimated values unbiased:
The "KaplanMeier" and "NelsonAalen" methods are estimators used in SurvivalDistribution. The distance of each point to its nearest neighbor point is censored by the distance of each point to the boundary of the observation region:
Compare the different edge correction methods:
Estimate the values of the nearest neighbor function with three different methods:
Applications (5)
The nearest neighbor function for spatially random data:
The empty space function for hardcore data:
Estimate the values of the empty space function with given data:
The mean distance a squirrel has to jump between trees in Lansing Woods can be computed using the nearest neighbor G function:
Mean distance between a typical point and its nearest neighbor (for positive support distribution, can be approximated via a Riemann sum of 1-CDF):
Compute the average minimum length of a cable needed to join a set of locations:
Analyze the probability of earthquakes depending on the existing data:
The probability of another earthquake within 10 miles from a historic earthquake location:
Properties & Relations (2)
NearestNeighborG is often compared with EmptySpaceF, which estimates the probability of finding another point within distance r from a reference point:
EmptySpaceF is a proper CDF:
Compare the estimates between NearestNeighborG and EmptySpaceF for point data generated by HardcorePointProcess:
The empty space and the nearest neighbor functions of a PoissonPointProcess are identical:
They both are equivalent to the CDF of an ExponentialDistribution:
Related Guides
History
Text
Wolfram Research (2020), NearestNeighborG, Wolfram Language function, https://reference.wolfram.com/language/ref/NearestNeighborG.html.
CMS
Wolfram Language. 2020. "NearestNeighborG." Wolfram Language & System Documentation Center. Wolfram Research. https://reference.wolfram.com/language/ref/NearestNeighborG.html.
APA
Wolfram Language. (2020). NearestNeighborG. Wolfram Language & System Documentation Center. Retrieved from https://reference.wolfram.com/language/ref/NearestNeighborG.html
BibTeX
@misc{reference.wolfram_2025_nearestneighborg, author="Wolfram Research", title="{NearestNeighborG}", year="2020", howpublished="\url{https://reference.wolfram.com/language/ref/NearestNeighborG.html}", note=[Accessed: 07-May-2026]}
BibLaTeX
@online{reference.wolfram_2025_nearestneighborg, organization={Wolfram Research}, title={NearestNeighborG}, year={2020}, url={https://reference.wolfram.com/language/ref/NearestNeighborG.html}, note=[Accessed: 07-May-2026]}