PairwiseDensityHistogram[{{y11,…,y1n},…,{ym1,…,ymn}}]
creates an array of density histograms by plotting the data columns against each other in pairs.
PairwiseDensityHistogram[data,bpsec]
plots density histograms with bins specified by bspec.




PairwiseDensityHistogram
PairwiseDensityHistogram[{{y11,…,y1n},…,{ym1,…,ymn}}]
creates an array of density histograms by plotting the data columns against each other in pairs.
PairwiseDensityHistogram[data,bpsec]
plots density histograms with bins specified by bspec.
Details and Options










- PairwiseDensityHistogram is a form of plot matrix.
- It plots high-dimensional data by creating a grid of individual plots of just two data columns at a time.
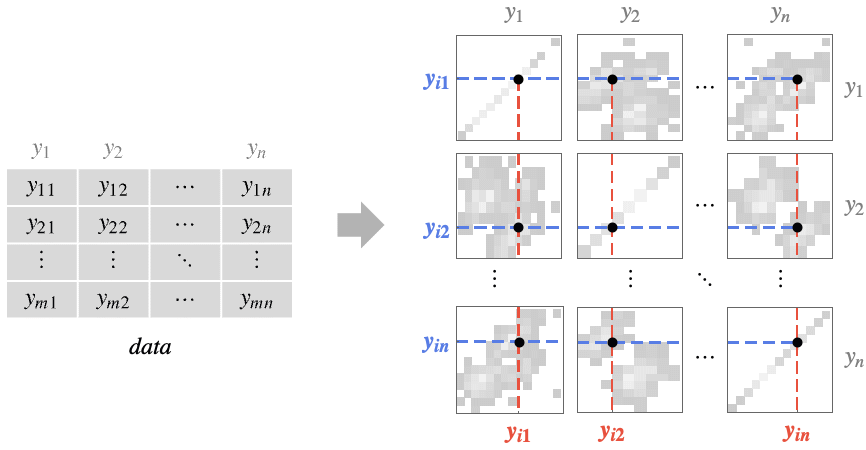
- The plot compares column yi={y1i,y2i,…,ymi} against column yj={y1j,y2j,…,ymj} by plotting a density histogram of the points {ykj,yki} in panel {i,j}.
- The panel at row
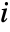 and column
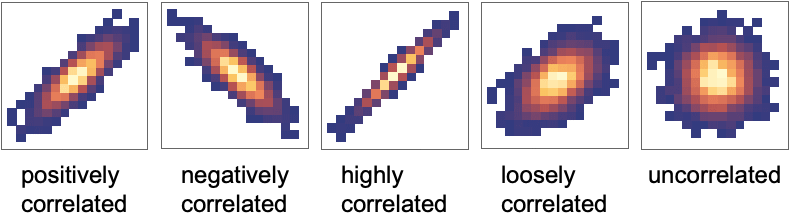
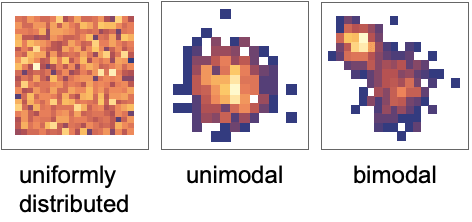
and column 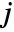 shows how the data columns yi and yj relate to each other. There are different types of features of the data that can be identified from those panels. Once the feature has been identified, it can also be quantified by further analysis of the data.
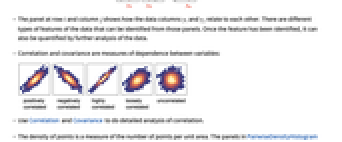
shows how the data columns yi and yj relate to each other. There are different types of features of the data that can be identified from those panels. Once the feature has been identified, it can also be quantified by further analysis of the data. - Correlation and covariance are measures of dependence between variables:
- Use Correlation and Covariance to do detailed analysis of correlation.
- The density of points is a measure of the number of points per unit area. The panels in PairwiseDensityHistogram effectively show the two-dimensional marginal densities:
- Use PairwiseDensityHistogram and PairwiseSmoothDensityHistogram to more directly visualize the densities.
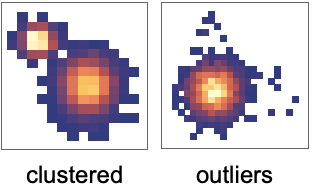
- Clusters are groups of similar points, indicating categories:
- The following bin specifications bspec can be given:
-
n use n bins {w} use bins of width w {min,max,w} use bins of width w from min to max {{b1,b2,…}} use bins [b1,b2),[b2,b3),… Automatic determine bin widths automatically "name" use a named binning method {"Log",bspec} apply binning bspec on log-transformed data fb apply fb to get an explicit bin specification {b1,b2,…} {spec1,spec2,…,specn} give different specifications for each data column yi - Possible named binning methods include:
-
"Sturges" compute the number of bins based on the length of data "Scott" asymptotically minimize the mean square error "FreedmanDiaconis" twice the interquartile range divided by the cube root of sample size "Knuth" balance likelihood and prior probability of a piecewise uniform model "Wand" one-level recursive approximate Wand binning - The function fb in PairwiseDensityHistogram[data,fb] is applied to a list of all {xi,yi} and should return an explicit bin list {{bx1,bx2,…},{by1,by2,…}}. In PairwiseDensityHistogram[data,{fx,fy}], fx is applied to the list of xi, and fy to the list of yi.
- PairwiseDensityHistogram[Tabular[…]cspec] extracts and plots values from the tabular object using the column specification cspec.
- The following forms of column specifications cspec are allowed for plotting tabular data:
-
{coly1,…,colyn} plot the colyi against colyj in a pairwise manner - PairwiseDensityHistogram takes the same options as Graphics, with the following changes and additions: [List of all options]
-
AspectRatio 1 ratio of height to width for each panel Background None background to use for each plot Frame Automatic whether to draw frames around each panel FrameTicks Automatic whether to label frame edges with ticks and labels GridLines Automatic whether to include gridlines in panels GridLinesStyle Automatic style for gridlines HeaderAlignment Center horizontal and vertical alignments of headers HeaderBackground Automatic background colors to use for headers HeaderDisplayFunction Automatic function to use to format headers Headers Automatic labels to use for each data column yi HeaderStyle None styles to use for headers PerformanceGoal $PerformanceGoal aspects of performance to try to optimize PlotHighlighting Automatic highlighting effect for points PlotLayout Automatic how to arrange the panels PlotLegends Automatic legends for data PlotMarkers None markers to use to indicate each point PlotStyle Automatic graphics directives to determine styles of points PlotTheme $PlotTheme overall theme for the plot Spacings Automatic horizontal and vertical spacings - PlotLayout can have the following settings:
-

"Descending" data columns going down and to the right 
"Ascending" data columns going up and to the right 
"DescendingHalfMatrix" lower half of the descending layout 
"AscendingHalfMatrix" upper half of the ascending layout - Headers specify the labels to use for each column in the data and will generally be displayed above each column and after each row in the final plot.
- Possible settings include:
-
None leave the plot columns and rows unlabeled Automatic automatically label columns and rows All always include column and row labels "Indexed" number the columns and rows 1, 2, …, n {lbl1,lbl2,…,lbln} use the given labels lbli - HeaderAlignment determines how data column labels are aligned with regard to the plot columns and rows.
- HeaderAlignment can take the following forms:
-
Center center the labels in the header positions {h,v} separate horizontal and vertical alignments within the header position {cols,rows} use col for plot columns and row for plot rows - HeaderBackground and HeaderStyle can take the following forms:
-
None use ambient styling sty use the style sty for all headers {sty1,sty2,…,styn} use the given styles styi for successive headers {cols,rows} use col for plot columns and row for plot rows - HeaderDisplayFunction determines how headers are displayed.
- Possible settings are:
-
Automatic automatic formatting None use unprocessed labels - The arguments to ColorFunction are yi1,yi2,…,yin. By default, the color function arguments are scaled per data column to be between 0 and 1.
- Use ColorFunctionScalingNone to use unscaled values or ColorFunctionScaling{cfsc1,cfsc2,…} to selectively scale column values.
- Possible settings for PlotHighlighting include:
-
Automatic automatically highlight positions in the panels None disable interactive highlighting -
 Highlight options with settings specific to PairwiseDensityHistogram
Highlight options with settings specific to PairwiseDensityHistogram
-
AlignmentPoint Center the default point in the graphic to align with AspectRatio 1 ratio of height to width for each panel Axes False whether to draw axes AxesLabel None axes labels AxesOrigin Automatic where axes should cross AxesStyle {} style specifications for the axes Background None background to use for each plot BaselinePosition Automatic how to align with a surrounding text baseline BaseStyle {} base style specifications for the graphic ContentSelectable Automatic whether to allow contents to be selected CoordinatesToolOptions Automatic detailed behavior of the coordinates tool Epilog {} primitives rendered after the main plot FormatType TraditionalForm the default format type for text Frame Automatic whether to draw frames around each panel FrameLabel None frame labels FrameStyle {} style specifications for the frame FrameTicks Automatic whether to label frame edges with ticks and labels FrameTicksStyle {} style specifications for frame ticks GridLines Automatic whether to include gridlines in panels GridLinesStyle Automatic style for gridlines HeaderAlignment Center horizontal and vertical alignments of headers HeaderBackground Automatic background colors to use for headers HeaderDisplayFunction Automatic function to use to format headers Headers Automatic labels to use for each data column yi HeaderStyle None styles to use for headers ImageMargins 0. the margins to leave around the graphic ImagePadding All what extra padding to allow for labels etc. ImageSize Automatic the absolute size at which to render the graphic LabelStyle {} style specifications for labels Method Automatic details of graphics methods to use PerformanceGoal $PerformanceGoal aspects of performance to try to optimize PlotHighlighting Automatic highlighting effect for points PlotLabel None an overall label for the plot PlotLayout Automatic how to arrange the panels PlotLegends Automatic legends for data PlotMarkers None markers to use to indicate each point PlotRange All range of values to include PlotRangeClipping False whether to clip at the plot range PlotRangePadding Automatic how much to pad the range of values PlotRegion Automatic the final display region to be filled PlotStyle Automatic graphics directives to determine styles of points PlotTheme $PlotTheme overall theme for the plot PreserveImageOptions Automatic whether to preserve image options when displaying new versions of the same graphic Prolog {} primitives rendered before the main plot RotateLabel True whether to rotate y labels on the frame Spacings Automatic horizontal and vertical spacings Ticks Automatic axes ticks TicksStyle {} style specifications for axes ticks
List of all options
Examples
open all close allBasic Examples (3)
Scope (4)
Options (44)
AspectRatio (2)
By default, PairwiseDensityHistogram uses an equal height-to-width ratio:
AxesStyle (4)
Background (2)
ChartBaseStyle (3)
Use ChartBaseStyle to style bins:
ChartBaseStyle combines with ColorFunction:
ColorFunction may override settings for ChartBaseStyle:
ChartLegends (4)
Legends use the same styles as the plot:
Use Placed to specify the legend placement:
Place the legend inside the plot:
Use LegendFunction to change the legend appearance:
ColorFunction (3)
Frame (2)
On PairwiseDensityHistogram, a frame is drawn on each subplot by default:
GridLines (2)
By default, PairwiseDensityHistogram does not draw grid lines:
Specify a GridLinesStyle on the grid:
HeaderBackground (2)
Headers (3)
By default, PairwiseDensityHistogram does not label headers:
ImageSize (6)
Use named sizes such as Tiny, Small, Medium and Large:
Specify the width of the plot:
Specify the height of the plot:
Allow the width and height to be up to a certain size:
Specify the width and height for a graphic, padding with space if necessary:
Setting AspectRatioFull will fill the available space:
Use maximum sizes for the width and height:
Use ImageSizeFull to fill the available space in an object:
PlotLayout (3)
PlotTheme (2)
Related Guides
Text
Wolfram Research (2024), PairwiseDensityHistogram, Wolfram Language function, https://reference.wolfram.com/language/ref/PairwiseDensityHistogram.html (updated 2025).
CMS
Wolfram Language. 2024. "PairwiseDensityHistogram." Wolfram Language & System Documentation Center. Wolfram Research. Last Modified 2025. https://reference.wolfram.com/language/ref/PairwiseDensityHistogram.html.
APA
Wolfram Language. (2024). PairwiseDensityHistogram. Wolfram Language & System Documentation Center. Retrieved from https://reference.wolfram.com/language/ref/PairwiseDensityHistogram.html
BibTeX
@misc{reference.wolfram_2025_pairwisedensityhistogram, author="Wolfram Research", title="{PairwiseDensityHistogram}", year="2025", howpublished="\url{https://reference.wolfram.com/language/ref/PairwiseDensityHistogram.html}", note=[Accessed: 20-April-2026]}
BibLaTeX
@online{reference.wolfram_2025_pairwisedensityhistogram, organization={Wolfram Research}, title={PairwiseDensityHistogram}, year={2025}, url={https://reference.wolfram.com/language/ref/PairwiseDensityHistogram.html}, note=[Accessed: 20-April-2026]}